In labs from Baltimore to Nijmegen, the boundary between chemistry and cognition is quietly shifting. Minimal “protocells” built from lipids and proteins can now sense a chemical cue and reorganize themselves, the first twitch of something like decision-making. Protein circuits inside mammalian cells perform winner‑take‑all classifications, turning messy signals into a single, decisive action. And tiny brain organoids show the building blocks of learning, hinting at biological hardware that adapts on the fly. I still catch myself rereading these papers, because if you strip the hype away, the core result remains startling: matter we assemble is beginning to choose.
The Hidden Clues
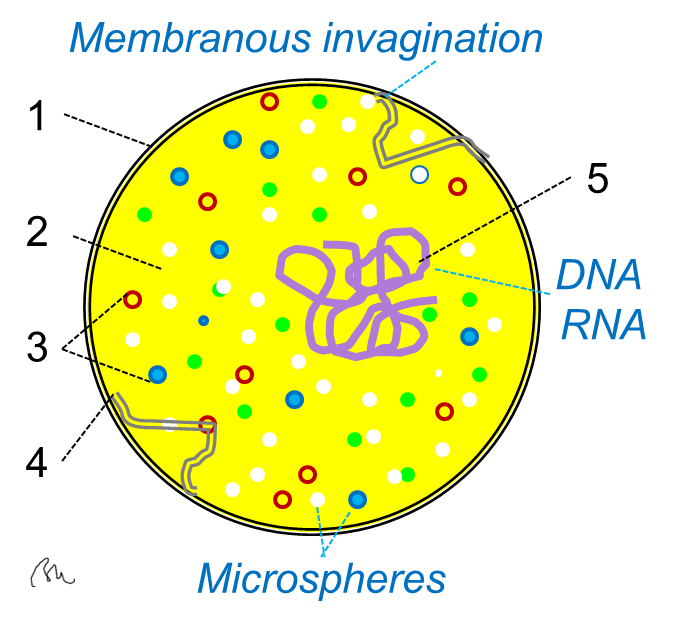
Scientists didn’t start by chasing “thinking cells.” They began with simpler questions: can an artificial membrane sense direction, can a reaction network remember, can a cell compute a choice without touching its genome. The answers arrived as subtle signatures – symmetry breaking in a synthetic bubble, a chemical reactor that predicts time series, a protein web that amplifies one signal while suppressing the rest.
At Johns Hopkins, a bare-bones protocell built from lipids and purified proteins sensed a gradient and reshaped itself, reproducing a key step cells use before they move. In parallel, a Nature study showed the old prebiotic formose reaction acting like a reservoir computer, classifying inputs and forecasting dynamics without transistors. Taken together, these are breadcrumbs on the path from physics to purpose.
From Ancient Tools to Modern Science

Our computers were born from sand heated into silicon; these new ones lean on chemistry that predated life. The formose reaction – long studied as a route to sugars on the early Earth – has so many feedback loops that it effectively computes when nudged with the right inputs. Researchers piped reagents into a stirred tank, read hundreds of molecules by mass spectrometry, and trained a simple linear model to extract the answer from the slurry’s evolving composition.
That might sound Rube Goldberg, but it demonstrates a principle with teeth: information can ride emergent dynamics, not just etched circuits. In a world hungry for low‑energy computation, a beaker that both stores memory and predicts the future deserves a seat at the table.
Inside the Molecular Machinery
Inside living cells, researchers are wiring up decisions using proteins instead of DNA edits. One landmark system – nicknamed a protein perceptron – balances self‑activation and mutual inhibition so that, when competing inputs arrive, only the strongest wins. Link that “win” to a functional outcome like apoptosis, and you’ve built a programmable reflex inside a mammalian cell.
Complementary work pushes DNA circuits and logic gates toward faster, more flexible molecular computing, while protocell engineers add sensors and cytoskeletal mimics to membrane bubbles. The theme is consistent: push computation down from the organism to the molecule, where decisions can be compact, analog, and frugal.
The Breakthrough Experiments
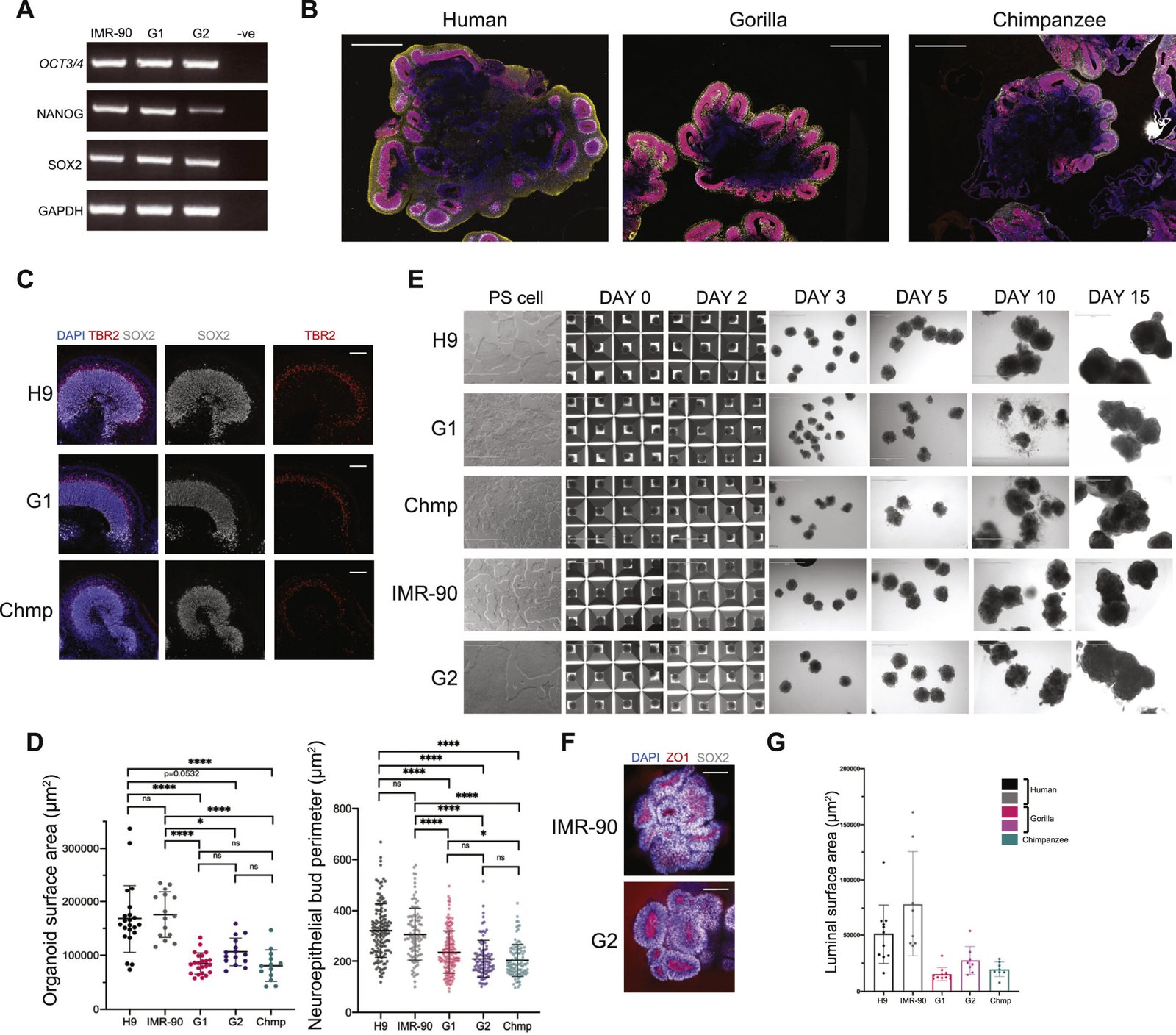
Two strands of evidence make the “thinking cell” headline feel less like a metaphor. First, human brain organoids – tiny clusters of neurons – now show the hallmarks of learning, including changes in connectivity and gene expression after controlled stimulation. These are not miniature minds, but they do exhibit plasticity that lets them adapt to incoming patterns.
Second, organoids can be harnessed as computing elements: research has shown that cultured neural networks can be trained to classify tactile stimuli patterns, including work on Braille character recognition. Meanwhile, self‑assembled “anthrobots” made from human tracheal cells crawled across damaged neural tissue in vitro and encouraged repair, an unexpected display of collective problem‑solving. None of these systems are conscious, but all of them act as if they can weigh inputs and alter behavior.
Why It Matters

Traditional chips are astonishing at arithmetic and large‑scale inference, but they gulp energy and struggle with noisy, embodied signals. Artificial cells flip the script: they live in noise. A protein circuit that sorts signals inside a T cell or a protocell that orients toward a chemical gradient could bring computation to the edge – inside tissue, in groundwater, on a leaf – without cloud connections or kilojoules to spare.
Compared with decades of gene‑circuit work, today’s protein‑level classifiers and gradient‑sensing protocells are faster, less brittle, and inherently biocompatible. Think diagnostics that only trigger when a disease signature dominates, or drug couriers that steer by the chemistry of inflammation. The payoff isn’t replacing GPUs; it’s marrying digital intelligence with living context.
Safety and Ethics
“Thinking” is a loaded word, and the risk isn’t rogue life – it’s misinterpretation. These systems compute; they don’t feel. Teams working on organoid intelligence are building ethical frameworks in lockstep with technical advances, emphasizing limits on complexity, monitoring for signatures of sentience, and ensuring strict oversight for any closed‑loop training.
On the biosafety front, most constructs are short‑lived, require special media, or lack the genes to survive outside lab conditions. That’s by design. The community has a chance to set norms early: transparency, risk assessments, red‑team testing, and public engagement before deployment.
The Future Landscape
Look one to five years out and three trends stand out. First, bottom‑up synthetic life projects aim to cross the Darwinian threshold – cells that grow, divide, and evolve – raising both opportunity and scrutiny. Second, lab groups are scaling the “training environments” for organoids, using closed‑loop stimulation and even automated protocol design to probe learning with the rigor of machine learning benchmarks.
Third, hybrid bio‑silicon architectures will move from demos to tools, with chemical reservoirs or organoids front‑ending sensors while silicon handles orchestration. In short, the map expands from petri dishes to platforms – and the winners will be the labs that pair caution with creativity.
What You Can Do Now

If this field excites you, you don’t need a wet lab to contribute. Support open data and preregistration for organoid and protocell studies, which helps separate robust results from fragile ones. Back interdisciplinary teaching – biochemistry for coders, control theory for biologists – because the next breakthroughs will come from people who speak both languages.
As a reader, cultivate nuance: celebrate computation in cells without inflating it into consciousness. If you’re in policy or philanthropy, invest in safety toolkits and community labs that practice responsible biology. The story is just beginning, and it will move faster – and safer – if more of us are paying attention.

Suhail Ahmed is a passionate digital professional and nature enthusiast with over 8 years of experience in content strategy, SEO, web development, and digital operations. Alongside his freelance journey, Suhail actively contributes to nature and wildlife platforms like Discover Wildlife, where he channels his curiosity for the planet into engaging, educational storytelling.
With a strong background in managing digital ecosystems — from ecommerce stores and WordPress websites to social media and automation — Suhail merges technical precision with creative insight. His content reflects a rare balance: SEO-friendly yet deeply human, data-informed yet emotionally resonant.
Driven by a love for discovery and storytelling, Suhail believes in using digital platforms to amplify causes that matter — especially those protecting Earth’s biodiversity and inspiring sustainable living. Whether he’s managing online projects or crafting wildlife content, his goal remains the same: to inform, inspire, and leave a positive digital footprint.